(And this post spawned a “SISTER POST” of sorts. Enjoy.)
UPDATED MEDIA
TIMELINE CHAPTERS
Here I want to offer a somewhat short refutation [NOT] of the perpetual myth about human and chimpanzee DNA being 99% similar. One friend included it in a comment to me:
- A cat shares 85 percent of our DNA along with dogs. Plants 15-20 percent . We share 90% of the genome with a banana. Chimpanzees 99% nearly…
Here is my short response:
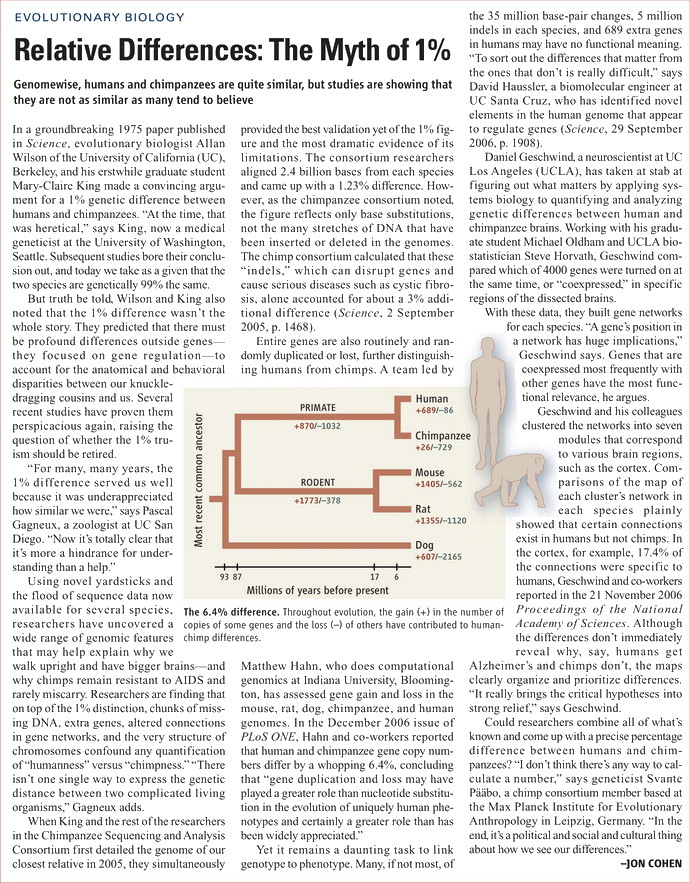
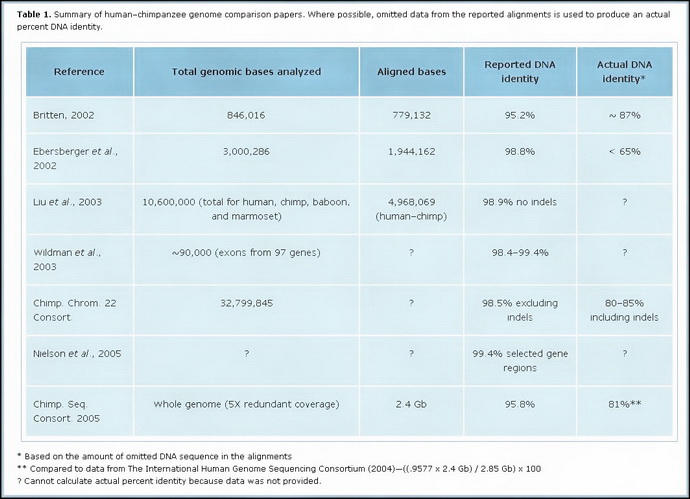
Not only that, but your idea of 99% is not a real stat as well. Many things have changed since that 1975 claim.* One example is that junk DNA is roundly refuted, and 2001 and 2005 Nature and Science Journal articles make clear that we share from 81% to 87% of DNA with chimps. That shouldn’t be a surprise since we both have eyes to see, stomachs to digest food, etc. So again, when I see you make claims above, rarely are they rooted in anything either current or true. * (CREATION.COM) The original 1% claim goes back to 1975.2 This was a long time before a direct comparison of the individual ‘letters’ (base pairs) of human and chimp DNA was possible—the first draft of the human DNA was not published until 2001 and for the chimp it was 2005. The 1975 figure came from crude comparisons of very limited stretches of human and chimp DNA that had been pre-selected for similarity. The chimp and human DNA strands were then checked for how much they stuck to each other—a method called DNA hybridization. (2. Cohen, J., Relative differences: the myth of 1%, Science 316(5833):1836, 2007; doi: 10.1126/science.316.5833.1836)
Even a recent 2006 TIME article continues the mantra when they say, “Scientists figured out decades ago that chimps are our nearest evolutionary cousins, roughly 98% to 99% identical to humans at the genetic level.” So while science moves on and corrects itself, our culture is stuck in what was said to be a proof, and reject what ACTUALLY an evidence against the evolutionary proposition. Similar refutations of evolutionary positions that Richard Dawkins and “Junk DNA.”
What do I mean by that? I mean that if something is said to be evidence and is used to promote [FOR] the evolutionary paradigm… and then it is shown not to be the case… wouldn’t it then logically be an evidence AGAINST this said paradigm? I think so.
MOVING ON… SORTA
Before zeroing in on the Chimp issue, one other quick note regarding a recent discovery that undermines this “similarity” idea. That is this study:
PJ MEDIA notes: A study published in the journal Human Evolution is causing quite the stir. In the words of Phys.org, “The study’s most startling result, perhaps, is that nine out of 10 species on Earth today, including humans, came into being 100,000 to 200,000 years ago.” So startling, in fact, that according to David Thaler, one of the lead authors of the study, “This conclusion is very surprising, and I fought against it as hard as I could.” The study’s very own author was so disturbed by how the conclusions challenged current scientific dogma that he “fought against it as hard as [he] could.” His “fight” gives credence to the study’s conclusions. His eventual acceptance, not to mention publication, of the conclusions speaks well of Thaler’s commitment to being a scientist first and an ideologue second. [….] This is no small matter for evolutionists because, as World Magazine helpfully summarizes: According to traditional evolutionary thinking, all living things on Earth share common ancestry, with species evolving through a slow process of random mutation, natural selection, and adaptation over roughly 3.8 billion years. The idea that humans and most animals suddenly appeared at the same time a mere 200,000 years ago or less does not fit with that model. (See more from my post, “Major DNA Study Undermines Evolution ‘In A Big Way’“) Obviously we differ on time-scales… but it sure seems like they are getting closer to mine over said time. But if one wishes to keep it ecumenical, here is a quote I love: >> J Warner Wallace, God’s Crime Scene: A Cold-case Detective Examines the Evidence for a Divinely Created Universe (Colorado Springs, CO: David C. Cook, 2015), 37.
Okay, back to the refutation of the 99% similarity. Here, Dr. Thomas Seiler, Ph.D., Physics, Technical University of Munich refutes compellingly this outdated TIME magazine article… and my friend:
…Most of you may have heard the statement that chimpanzees and humans are having 99% of their genes in common. However, what you are usually not told is that this result was not based on comparing the entire DNA of man and ape but only on comparing a very small fraction of it (ca. 3 %). The function of the other 97% of the genetic code was not understood. Therefore, it was concluded that this DNA had no function at all and it was considered “leftover junk from evolution” and not taken into consideration for the comparison between man and ape. Meanwhile, modern genetics has demonstrated for almost the entire DNA that there is functionality in every genetic letter. And this has led to the collapse of the claim that man and chimpanzee have 99% of their DNA in common. In 2007, the leading scientific journal Science therefore called the suggested 1% difference “a myth.” And from a publication in Nature in 2010 comparing the genes of our so-called Y-chromosome with those of the chimpanzee Y-chromosome we know now that 60% of human Y-chromosome is not contained in that of the chimpanzee. This represents a difference of one billion genetic letters, known as nucleotides. And modern genetics has recently made another important discovery which was very unexpected. Researchers found that all of the different groups of humans on earth, wherever they live and whatever they look like, have 99.9% of their genes in common. This leads to a problem for the hypothesis of evolution because if humans really were descended from the apes, then how could it be that we only have 40% of our Y-chromosome in common with the apes but at the same time there is almost a complete genetic identity among all humans? If there had been an evolution from ape to man then it should still go on among men and reveal significant genetic differences. These recent discoveries therefore drastically widen the gap between man and the animals. And they confirm that there are in reality no such things as human “races”. Asians, Europeans, Africans and Indigenous people from America and Australia only have superficial differences like color of skin or shape of the nose but they are all extremely similar on the genetic level. And these recent breakthrough discoveries even go further. Today, because of the extreme similarity of the human genome, it is considered a well-established fact among geneticists, that all humans living on earth now are descended from one single man and from one single woman. In order to convince yourself of this you only have to search in the internet for the terms “mitochondrial Eve” or “Y-chromosome Adam”. These names were given by evolutionists in an ironic sense but now many regret that choice of name because this discovery perfectly confirms the Catholic Doctrine of Creation which has taught for 2000 years that all humans are brothers and sisters descended from one single human couple, the real historical persons Adam and Eve, not from a multitude of subhuman primates…. (Via LIFE SITE NEWS)
Here is a visual of the varying studies (click to enlarge in another window):
This video evaluates the claim that humans and chimps have 98% to 99% DNA similarity.
DR. JONATHAN SARFATI passed this on to me in conversation (click to enlarge as PDF):
Wow. Enough said? Or will this myth still infect the brains of people wishing something to be true that continue to lose evidences for? One other noteworthy exchange from that conversation I wish to note here.
THE BELOW IS DATED MAY 2010
See also: Pseudogenes Predicted To Fail by Creationists (Vestigial Response Added)
Bottom Line:
The failure to recognize the full implications of [non-protein–coding DNA] may well go down as one of the biggest mistakes in the history of molecular biology. (cited in: Gibbs, W.W., The unseen genome: gems among the junk, Scientific American 289[5]:26–33, November 2003.)
…even the staunchest critics of creation theory recognize that “[i]t is impossible to prove absence of function for any region of DNA.” Edward E. Max, “Plagiarized Errors and Molecular Genetics,” Sec. 5.4 . From: A Critique of “29 Evidences for Macroevolution”
Talk Origins in its 29+ Evidences for Macroevolution (written between 199 and 2004) quotes a 1999 study that purportedly proves their stance:
Furthermore, all of these genes have accumulated mutations at the exact rate predicted (the background rate of mutation for neutral DNA regions like pseudogenes) (Ohta and Nishikimi 1999).
Well, the tables have turned on neo-Darminian claims yet once again and science (not “scientism”) advances:
- The slow, painful death of junk DNA
- Potentially decisive evidence against pseudogene “shared mistakes”
- GULO pseudogenes have been intensively investigated
- Prediction 6: Anatomical Vestigial Structures
- Prediction 17: Functional Molecular Evidence — Protein Functional Redundancy: A Critique Of “29 Evidences For Macroevolution” Part 4
- Dr. Edward Max on Plagiarism & Shared Genetic Mistakes
- Design inferences and the human inability to produce Vitamin C
- Adam and Eve, Vitamin C, and Pseudogenes

It is worth mentioning that this supposed truth for evolutionary theory is further along the wrong prediction mark than the proof of “Junk DNA” was along this path when I was debating in 97′ with people over at the Discover site. (In fact, this “proof” is connected to the “junk DNA” position.) In 1997 people — in debate situation — said that evolution predicts junk DNA and that they have known of junk DNA for quite some time, thus proving evolutionary explanatory power. IN 2003, one of the first articles to hit the mainstream scientific press that this long held assumption was in jeopardy hit Discover and Scientific American magazines, two subscriptions I receive out of the many mags I get. Pictured is the first Discover magazine (yes, I have kept them) ruminations that this “proof” was disintegrating, like the vestigial organs “proof” did, cytochrome C differences, bacterial evolution, etc.
So debating a topic like this supposed proof is funny because it is much further along the scientific path than it was when I first debated the issue. Not to mention all the “evolutionary tree” models that are supposed to be factual when in fact their order is a topic of much debate. For instance, I posted a bit earlier a layman’s article on the argument within the evolutionary camp about mankind coming from orangutans rather than a chimpanzee split in the family tree… (this earlier post mentions the belief within the evolutionary camp that man came from some aquatic ancestor rather than orangutans). A few people likewise believe that apes (Gorillas) are descended from mankind in some way. For instance, Dr. Aaron G. Filler:
Dr. Aaron G. Filler, M.D., Ph.D. studied evolutionary theory under some of the leading biologists and anthropologists of our time: Ernst Mayr, Stephen J. Gould, David Pilbeam, and Irven DeVore. A neurosurgeon at the Institute for Spinal Disorders at Cedars Sinai Medical Center and past associate director of the Comprehensive Spine Center at UCLA, Dr. Filler has been a leading innovator in medical imaging and neuroscience. He is the author of Do You Really Need Back Surgery? (Oxford University Press), as well as numerous scientific articles and patents.
He wrote a book entitled The Upright Ape: A New Origin of the Species, in which he follows the footsteps of another well respected scientist showing that apes are most likely a breakaway group from mankind. The other scientist I mention is Dr. Geoffrey H. Bourne:
He was Director of Yerkes Regional Primate Research Center at Emory University, England. Dr. Bourne is Oxford educated, and is an American cell biologist/anatomist who was considered by most to be the worlds leading primatologist.
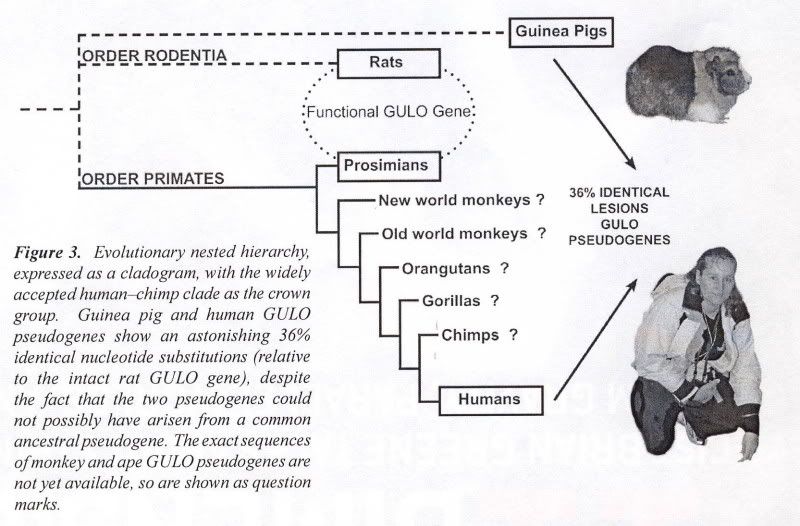
He said that apes are descended from man. Why would men of science believe such a thing? Because science has never seen any information being added to the evolutionary upward “slant” that is required by its theory (Darwinism, e.g., “scientism”). So since apes are less than us, Dr. Bourne says that science [not “scientism”] proves his theory due to observable facts. (SEE APPENDIX) All this though, junk DNA being disproved and a differing order of our family tree are other impediments to the Vitamin C Pseudogene argument. In order for this argument to work (and the subsequent predictions made to be coherent), he [the evolutionist] has to have all his ducks in order, this is something he does not have the luxury of (see image at top). One of the papers linked above (GULO pseudogenes have been intensively investigated) and its in-depth research led it two authors to say the following:
When examined in detail, the full pseudogene dataset we collected does not lend itself to a reasonable neo-Darwinian interpretation. Using standard bioinformatics tools and principles, we present alternative designs for at least the exon X portion of the GULO gene. These may be plausible due to nucleotide patterns being relevant as regulatory signals or the favouring of some codons for various possible reasons. We do accept that some mutations have occurred in this exon. But these novel proposals imply that the ancestors of the organisms studied may well never have had the exact same GULO sequence.
That last sentence really shows the assumptions made in this “proof” that steps away from true science and into the realm of historical science. And the historical sciences are fraught with bickering, changing branches of human lineage, and naturalistic philosophies that make the vitamin C pseudogene argument pretty vacuous.
The following is a good round-up of the problems seen in this line of argument, by Sean D. Pitman MD:
…in 2003, the same Japanese group published the complete sequence of the guinea pig GLO pseudogene, which is thought to have evolved independently, and compared it to that of humans [Inai et al, 2003]. 21 Surprisingly, they reported many shared mutations (deletions and substitutions) present in both humans and guinea pigs. Remember now that humans and guinea pigs are thought to have diverged at the time of the common ancestor with rodents. Therefore, a mutational difference between a guinea pig and a rat should not be shared by humans with better than random odds. But, this was not what was observed. Many mutational differences were shared by humans, including the one at position 97. According to Inai et al, this indicated some form of non-random bias that was independent of common descent or evolutionary ancestry. The probability of the same substitutions in both humans and guinea pigs occurring at the observed number of positions was calculated, by Inai et al, to be 1.84×10-12 – consistent with mutational hotspots. What is interesting here is that the mutational hot spots found in guinea pigs and humans exactly match the mutations that set humans and primates apart from the rat (see figure below). 21,22 This particular feature has given rise to the obvious argument that Inai et al got it wrong. Reed Cartwright, a population geneticist, has noted a methodological flaw in the Inai paper: “However, the sections quoted from Inai et al. (2003) suffer from a major methodological error; they failed to consider that substitutions could have occurred in the rat lineage after the splits from the other two. The researchers actually clustered substitutions that are specific to the rat lineage with separate substitutions shared by guinea pigs and humans. . . If I performed the same analysis as Inai et al. (2003), I would conclude that there are ten positions where humans and guinea pigs experienced separate substitutions of the same nucleotide, otherwise known as shared, derived traits. These positions are 1, 22, 31, 58, 79, 81, 97, 100, 109, 157. However, most of these are shown to be substitutions in the rat lineage when we look at larger samples of species. When we look at this larger data table, only one position of the ten, 81, stands out as a possible case of a shared derived trait, one position, 97, is inconclusive, and the other eight positions are more than likely shared ancestral sites. With this additional phylogenetic information, I have shown that the “hot spots” Inai et al. (2003) found are not well supported.” (see Link) It does indeed seems like a number of the sequence differences noted by Cartwright are fairly unique to the rat – especially when one includes several other species in the comparison. However, I do have a question regarding this point. It seems to me that there simply are too many loci where the rat is the only odd sequence out in Exon X (i.e., there are seven and arguably eight of these loci). Given the published estimate on mutation rates (Drake) of about 2 x 10-10 per loci per generation, one should expect to see only 1 or 2 mutations in the 164 nucleotide exon in question (Exon X) over the course of the assumed time of some 30 Ma (million years). Therefore, the argument of the mutational differences being due to mutations in the rat lineage pre-supposes a much greater mutation rate in the rat than in the guinea pig. The same thing is true if one compares the rat with the mouse (i.e., the rat’s evident mutation rate is much higher than that of the mouse). This is especially interesting since many of the DNA mutations are synonymous. Why should essentially neutral mutations become fixed to a much greater extent in the rat gene pool as compared to the other gene pools? Wouldn’t this significant mutation rate difference, by itself, seem to suggest a mutationally “hot” region – at least in the rat? Beyond this, several loci differences are not exclusive to the rat/mouse gene pools and therefore suggest mutational hotspots beyond the general overall “hotness” or propensity for mutations in this particular genetic sequence. Some have noted that although the shared mutations may be the result of hotspots, there are many more mutational differences between humans and rats/guinea pigs as compared to apes. Therefore, regardless of hotspots, humans and apes are clearly more closely related than are humans and rats/guinea pigs. The problem with this argument is that the rate at which mutations occur is related to the average generation time. Those creatures that have a shorter generation time have a correspondingly higher mutation rate over the same absolute period of time – like 100 years. Therefore, it is only to be expected that those creatures with relatively long generation times, like humans and apes, would have fewer mutational differences relative to each other over the same period of time relative to those creatures with much shorter generation times, like rats and guinea pigs. What is interesting about many of these mutational losses is that they often share the same mutational changes. It is at least reasonably plausible then that the GULO mutation could also be the result of a similar genetic instability that is shared by similar creatures (such as humans and the great apes). This same sort of thing is seen to a fairly significant degree in the GULO region. Many of the same regional mutations are shared between humans and guinea pigs. Consider the following illustration yet again: Why would both humans and guinea pigs share major deletions of exons I, V and VI as well as four stop codons if these mutations were truly random? In addition to this, a mutant group of Danish pigs have also been found to show a loss of GULO functionality. And, guess what, the key mutation in these pigs was a loss of a sizable portion of exon VIII. This loss also matches the loss of primate exon VIII. In addition, there is a frame shift in intron 8 which results in a loss of correct coding for exons 9-12. This also reflects a very similar loss in this region in primates (see Link). That’s quite a few key similarities that were clearly not the result of common ancestry for the GULO region. This seems to be very good evidence that many if not all of the mutations of the GULO region are indeed the result of similar genetic instabilities and that are prone to similar mutations – especially in similar animals. As an aside, many other genetic mutations that result in functional losses are known to commonly affect the same genetic loci in the same or similar manner outside of common descent. For example, achondroplasia is a spontaneous mutation in humans in about 85% of the cases. In humans achondroplasia is due to mutations in the FGFR2 gene. A remarkable observation on the FGFR2 gene is that the major part of the mutations are introduced at the same two spots (755 C->G and 755-757 CGC->TCT) independent of common descent. The short legs of the Dachshund are also due to the same mutation(s). The same allelic mutation has occurred in sheep as well. (SOURCE)

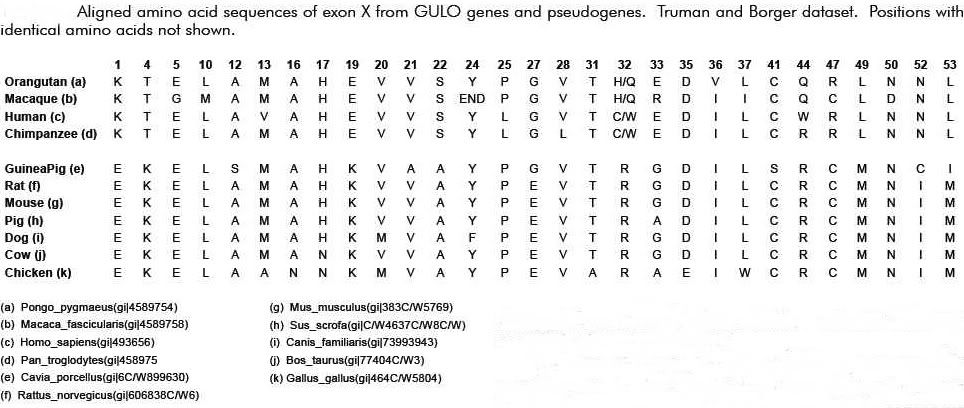
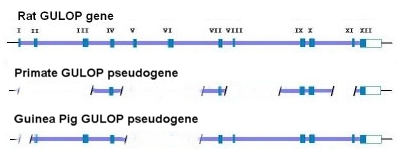
All this is to say, yet again, that this “evidence” is anything but!
By definition, pseudogenes lack a function. However, the classification of pseudogenes generally relies on computational analysis of genomic sequences using complex algorithms. This has led to the incorrect identification of pseudogenes. For example the functional, chimeric gene jingwei in Drosophila was once thought to be a processed pseudogene. It has been established that quite a few pseudogenes can go through the process of transcription, either if their own promoter is still intact or in some cases using the promoter of a nearby gene; this expression of pseudogenes also appears to be tissue-specific. In 2003, Hirotsune et al. identified a retrotransposed pseudogene whose transcript purportedly plays a trans-regulatory role in the expression of its homologous gene, Makorin1 (MKRN1) (see also RING finger domainubiquitin ligases), and suggested this as a general model under which pseudogenes may play an important biological role. Other researchers have since hypothesized similar roles for other pseudogenes. A bioinformatics analysis has shown that processed pseudogenes can be inserted into introns of annotated genes and be incorporated into alternatively spliced transcripts. Hirotsune’s report prompted two molecular biologists to carefully review scientific literature on the subject of pseudogenes. To the surprise of many, they found a number of examples in which pseudogenes play a role in gene regulation and expression, forcing Hirotsune’s group to rescind their claim that they were the first to identify pseudogene function. Furthermore, the original findings of Hirotsune et al. concerning Makorin1 have recently been strongly contested; thus, the possibility that some pseudogenes could have important biological functions was disputed. Additionally, University of Chicago and University of Cincinnati scientists reported in 2002 that a processed pseudogene called phosphoglycerate mutase 3 (PGAM3P) actually produces a functional protein and two 2008 publications in Nature discusses that some endogenous siRNAs are derived from pseudogenes, and thus some pseudogenes play a role in regulating protein-coding transcripts.
AIG & PDF Sources [PDF source near top]:
….Many of those mammals found unable to synthesize ascorbic acid have regions of their genome that are believed to correspond to parts of the functional GULO gene that is found in those mammals found capable to synthesizing GULO, and thus vitamin C. Evolutionists have cited these apparently vestigial remnants of GULO to make dysteleological arguments against an Intelligent Designer. In addition, they have argued that lesions found in common between the orthologous GULO pseudogenes of simian primates (‘shared mistakes’) argue strongly for their origins from a common ancestor, and all but rule out an independent inactivation of the GULO gene among different simian primates (including humans). Previous studies of the orthologous primate GULO gene and pseudogene have focused on those parts of a few exons that appear to correspond between humans and rats. A more recent study1 is much more comprehensive. It is now believed that, relative to the 12 exons that comprise the functional rat GULO gene, the human GULO pseudogene is limited to counterparts of exons 4, 7, 9, 10, and 12. Owing to the fact that the guinea pig and the simian primates are obviously not sister groups [see figure near top], it is impossible for the guinea pig GULO pseudogene and the human GULO pseudogene to have both originated from the same ancestral pseudogene. Furthermore, not only are the inactivations of GULO in the guinea pig and primates clearly independent events based on phylogenetic analysis [see figure near top], but also on inferred evolutionistically believed times of inactivation…. Inai, Y., Ohta, Y. and Nishikimi, M., The whole structure of the human non-functional L-gulono-γ-lactone oxidase gene—the gene responsible for scurvy—and the evolution of repetitive sequences thereon, J. Nutritional Science and Vitaminology (Tokyo)49(5):315–319, 2003.
UPDATE:
The “pseudogene” argument (of which Vitamin C Pseudogene is part of) is being shown to not be a “vestige” as claimed and “predicted” by Darwinian theory. Much like the Haeckel’s list of over 100 vestigial organs, science shows that predictions made like this fail. If the predictions of design were assumed, more lives would have been saved and true science would have flourished. JBS Haldane however, said (predicted) that “natural selection cannot possibly select for millions of new mutations over the course of human evolution, Kimura developed the idea of “Neutral Evolution.” If “Haldane’s Dilemma” was correct, the majority of DNA must be non-functional. Hence, the pseudogene argument. This is an argument that has failed due to recent science, no predictive power left in it. Which is no small issue, since, it is this “predictive value” that makes this argument valid. Faulkner (in 2009) worked with mice and humans and showed clearly that this RNA/Retrotranposons relationship were not random, and thus transcription is shown to be influenced wholly or partly by this “junk DNA.” — “Therefore, more than one third of the mouse and human genomes, previously thought to be non-functional, may play some role in the regulation of gene expression” (scientists from Jackson Maine, USA laboratories). Creationists have long figured this “junk DNA” to be functional… making design predictions more accurate. Thankfully, not everyone bought this idea. In the late 1980s, New Zealand–born Australian immunologist Malcolm Simons recognized patterns, or order, in the non-coding DNA that indicated to him that the code must have a function, but others ridiculed the idea. In the mid-1990s, he patented the non-coding DNA (95%) of all organisms on Earth. The company he founded, Genetic Technologies, now reaps license fees from all technologies being developed to cure disease that involve the non-coding DNA. It’s quite controversial, of course, paying such license fees. And since factors involved in all sorts of diseases, such as breast cancer, Crohn’s disease, Alzheimer’s, heart disease, ovarian and skin cancer, are being found in the ‘junk’, Genetic Technologies is doing quite well. The “vitamin C gene” in question here (L-gulano-g-lactone oxidase gene) is only a valid argument if: a) a non-functional version of the gene was shown to be functional at some point in our human lineage, it says nothing about ancestors (kinds, or species) who could have been created with an active gene. b) If the lineage of man can truly be traced through evolutionary ancestors, which it cannot. c) The same gene could have been inactivated by the same mutation occurring independently is said to be too improbable by strict neo-Darwinists. (Ironic) However, if there is, or was, a mechanism of mutation that favors certain locations in the gene (a hot spot), the odds against an independent occurrence of the mutation drop according to the strength of that bias. This, by-the-way, is more probable considering the fall of the pseudogene argument. In other words, this prediction would now be of a higher order and is an example of loss of function and a disordering of optimum ability. Something the creation model predicts, it is the opposite of what Darwinism requires. d) Likewise, in order for this line of reasoning to work, a human genome must be compared to an organism with a functioning gene for synthesizing vitamin C. Of course we know they found it in rats (Nishikimi, 1994). Four of the 12 exons (Science Dictionary: A segment of a gene that contains information used in coding for protein synthesis; exons are the modular coding regions of the gene). Here is the downfall, two-thirds of the homologous rat gene is completely missing. Which is to say: “This DNA sequence, labeled as a pseudogene, might have an entirely different function than the rat gene.” This would be — with recent peer reviewed articles in Discover, Scientific American, Nature, Science Weekly, and other pubvlications — be of a higher order than merely saying it is a pseudogene. In other words, the previous predictions are mute and shows that the argument is from silence, or non-science. Stating that only the last enzyme is missing for the pathway to convert glucose to vitamin C might imply to the untrained individual that there is a biochemical pathway that leads to a dead end. Actually, the biochemical pathway that leads to the synthesis of vitamin C in rats also leads to the formation of five-carbon sugars in the pentose phosphate pathway present in virtually all animals (Linster and Van Schaftingen 2007). There are several metabolic intermediates in this pathway illustrating that these substances can be used as precursors for many compounds in the cell. In the pentose phosphate pathway, five-carbon sugars are made from glucose (a six-carbon sugar) to be used in the synthesis of DNA, RNA, and many energy producing substances such as ATP and NADPH (Garrett 1999). Animals that synthesize vitamin C can use both pathways illustrated in the simplified diagram below. Humans and the other animals “less fortunate” than rats only use the pentose phosphate pathway. There is no dead end or wasted metabolic intermediates, and there is no need to have the enzyme to make vitamin C since humans are able to get all of the vitamin C they need from food substances. Thousands of human pseudogenes have been catalogued, but in spite of the similarities to functional genes, the exact role of pseudogene sequences in the genome are not known by any scientist. It is not necessary to assume that pseudogenes are remnants of once functioning genes that have been lost and now clutter the genome like junk in a rubbish heap. Criswell, D. 2007. Adam and Eve, Vitamin C, and Pseudogenes. Acts & Facts. 36 (5).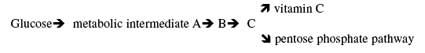
Conclusion Then
As was said before the known factors of the pentose phosphate pathways:
- The failure to recognize the full implications of [non-protein–coding DNA] may well go down as one of the biggest mistakes in the history of molecular biology. (cited in: Gibbs, W.W., The unseen genome: gems among the junk, Scientific American 289[5]:26–33, November 2003.)
- …even the staunchest critics of creation theory recognize that “[i]t is impossible to prove absence of function for any region of DNA.” (Edward E. Max, “Plagiarized Errors and Molecular Genetics,” Sec. 5.4 . From: A Critique of “29 Evidences for Macroevolution”)
So, to sum up, at best this could be viewed as a theological argument: “what should have God done if he did create ‘A’.” At worst it is an argument that has not evolved with science correcting itself and is stuck in the 90’s science and not the mid-to-late part of the first decade of the beginning of the 21st century — science. Science has evolved, have the evolutionists?
This is an excerpt from the much hated book, Of Pandas and People, pp 34-40 (click to enlarge):
Here is the continuation of the devolving of primates:
APPENDIX
1. It takes 6 million years for one mutation to arise in a DNA binding site.
2. 6 million years is how long, historically, one has to get from primates to humans.
3. Humans differ from primates in hundreds of ways, requiring probably thousands of different gene mutations.
4. What’s more, many of the differences between primates and humans would require coordinated mutations. For example, the correct legs, feet, pelvis, spine, and neck to walk upright are differences which individually are useless by themselves.
5. Experiments with bacteria (i.e. with population sizes and mutation rates much higher than primates and humans) suggest that a single protein cannot mutate six or more times in a coordinated fashion in a timespan less than the known age of the universe.
6. Given these numbers, it is impossible for humans to have evolved from primates.